In a eukaryotic cell, the length of the DNA molecule is approx. 1m (approx.6ft). In the nucleus, DNA is organized in a hierarchical order with the help of proteins collaborating to create a structure called chromatin. The Major The nucleosome is a kind of chromatin unit made up of DNA and histone protein. Histone protein gives structural support to a chromosome [1]. Histone proteins, lapping around the DNA is an octamer whose modification by acetylation after translation leads to activation of the transcription process regulated by the enzyme histone acetyltransferase (HAT) [2]. Acetylation occurs in the lysine residues of N-terminal tails of histones. HAT transfers the acetyl group present in the acetyl-CoA to the lysine residue containing the ɛ-NH3+ group at the N-terminal tails [3]. Due to the acetylation, the interaction between DNA and histone protein varies. The reverse process of acetylation is deacetylation which is catalysed by the enzyme HDAC. The mechanism of HDAC is either NAD+ dependent or NAD+ independent [4]. HDAC removes the acetyl group from the lysine residue which recondensed the DNA. NAD+-dependent HDAC can be classified into three classes according to their similarity to the yeast proteins RPD3 (class 1), HDAC1 (class 2) and SIRT2 (class 4) whereas Zn-dependent HDAC is HDAC11 [5]. Due to this acetylation-deacetylation process in histone modification, the proliferation and cell differentiation occur which causes various kinds of diseases such as several types of tumours, prostate cancer, colon cancer, breast cancer, skin cancer, cardiological disease, neurological diseases, vascular disease, genetic disorders related disease Rubinstein-Taybi Syndrome, Fragile X syndrome and it also influences the metabolism process [6-12]. HDAC inhibitors and HAT inhibitors play an important role in the treatment of diseases caused by histone modifications [12-13].
In this review, we have discussed the histone protein and its role in DNA binding. Then the purpose of the epigenetic modification and its result will also be explained. The process of epigenetic modification, acetylation and deacetylation, and the enzymes involved in this process are HAT and HDAC respectively will be covered in more detail. Then we’ll focus on the diseases associated with epigenetic modification. After that, the classes of inhibitors, their activity and their uses in the current clinical research for drug development will be discussed.
Histone Protein and post-translational modification
Histone has four types- H2A, H2B, H3, H4. These all subunits are single monomers, but these subunits are joined twice to create an octameric core with 146-147 base pairs of DNA firmly wrapped around it. The variable length of linker DNA separates nucleosome cores, which are linked by the linker histone H1 [1].
DNA replication and gene expression both are controlled by the overall structure of chromatin and its domains. Variable length of N-terminal tails and histone domains are present in core histones. These N-terminal tails are abundant in lysine and arginine and all the modifications like acetylation may occur in the side chain of lysine, the peptide bond or peptide backbone modification does not take place [14]. The N-tail plays a vital role in extensive post-translational modifications (PTMs), such as acetylation and phosphorylation. Acetylation of histone protein has been controlled by the transcription, whereas acetylation of histone is associated with silencing. PTMs are also implicated in chromatin assembly and DNA replication. PTMs consist of the epigenome which comprises changes to DNA that are connected to proteins. These epigenetic modifications are switches for the regulation of gene expression and are chemical modifications of the DNA and histones that do not change the DNA sequence. Changing the chromatin structure is correlated with the phosphorylation of linker histone [15].
As a result, several recent studies contend that histones play an important role in the control of biological processes such as transcription. It is more essential than understanding histone biology on a structural basis.
Structure and Specificity in Gene Regulation
A 7 resolution crystallographic study with a (H3)2(H4)2 tetramer yielded the structure of the nucleosome core particle, with two H2A H2B dimers at the centre and DNA wrapped around it. [1]. The histone fold domain is found in each core histone protein, which is made up of three -helices linked by two loops. The handshake motif allows for heterodimeric connections between core histones. Each of the core histones has an N-terminal tail that binds to a wide range of PTMs. In general, the main histone protein genes are found in gene clusters that are largely expressed during the S phase of the cell cycle. These histones are involved in the packaging of freshly produced DNA. Some histone variations are expressed throughout the cell cycle and are not limited to the S phase of the cell cycle [16]. Now we will study the structure of subunits of histone proteins.
Histone H2A: The core histone is H2A which has the largest number of variants. H2AZ and H2AX are the variants of histone H2A, mainly found in most eukaryotes. Histone H2A is characterized by its genome localization and its divergent C-terminal sequences. H2AZ has been involved in transcriptional activation in yeast and preventing the spread of silent heterochromatin. H2AX histone variants are distinguished by a C-terminal extension containing the consensus SQ[E/D] sequence, which denotes a hydrophobic amino acid [1].
Histone H2B: H2B histone variations are rare, yet they play important roles in chromatin compaction during gametogenesis. Furthermore, H2B variations have been discovered in male gametic cells from lily and, more recently, in bovine and human spermatozoa, albeit their particular functions remain unknown.
Histone H3: H3.3, CenH3, and H3.4 are the components of the histone H3 variations. The H3.3 variant is present in transcriptionally active chromatin and is not S-phase regulated. The H3.4 component is only found in the testis. H3 histone variations are detected in primordial spermatocytes. CenH3 variants are located in centromeric chromatin, and their N-terminal tails are divergent, with little sequence resemblance to conventional CenH3. [1]
Histone H4: The most conserved histone is the histone H4 variation. In the nucleosome core particle, H4 histone interacts with the other three core histone proteins. There are even identical sequence variations that are expressed separately within a cell cycle, as contrast to the main synthesis period for histones in the S phase [17]
Histone Acetylation and Role of Enzyme
Allfrey et al. first reported histone acetylation in 1964 [18]. It occurs in the nucleus of the eukaryotic cells which is regulating gene transcription and expression, so it is an epigenetic modification. Basically, DNA wraps tightly to histone proteins, to lose this bind from histone acetylation take place because if DNA does not get free from protein, then it can’t further proceed towards gene regulation. The enzymes that help the acetylation process are known as histone acetyltransferases (HATs). Mainly the enzyme HATs acetylated lysine amino group from acetyl-CoA to form ε-N acetyl-lysine [19]. Histone is a positively charged protein and DNA is negatively charged, that’s why their interaction is so strong and tight. Here acetylation removes the positive charge from histone protein, and it becomes negative. Acetylation reduces the affinity of the histone for DNA, so their interaction decreases, and DNA loses its coiling pattern, as a result, the transcription process is activated. Acetylation is an activator of transcription. Acetylation is a reversible process, so histone deacetylases (HDACs) are the enzymes that counteract that activity.
Non-Histone Protein and Post Translational Modification
Those proteins which remain in chromatin after removed the histone proteins are known as non-histone proteins. A large group of heterogeneous non-histone proteins plays a role in the compaction and organization of higher-order chromosome structure, they are also involved in nucleosome remodeling, DNA replication, RNA synthesis and processing, nuclear transport, steroid hormone activity, and the transition from interphase to mitosis. Proteins that are not histones, such as scaffold protein, DNA polymerase, heterochromatin protein1, and polycomb are very common. This non-histone protein is acidic in nature [20].
Histone Acetyltransferase
A group of enzymes that transfers acetyl group from acetyl coA to lysine sidechain containing epsilon amino group is known as HAT. Depending upon the mechanism of catalysis HATs can be classified into two classes such as HAT A localised in nucleus, takes part in transferring acetyl group from acetyl coA to lysine on histone & HAT B localised in cytoplasm, takes part in transferring acetyl group from acetyl coA to lysine residue on newly synthesised free histone [2,21].
Classification and Role of Enzyme:
Based on subcellular localization HATs are classified into two classes, one types A HATs and another one is type B HATs [22] In the nucleus, type A HATs are found that are involved in the regulation of gene expression through acetylation. Base on their homology and acetylation mechanism the type A HATs are further classified. PCAF, Gnc5, and ELP3 are members of the GNAT family. The CBP/p300 family members are CBP and p300. Tip60, MOZ, MORF, HBO1, and HMOF all are members of the MYST family. TAF1 and TIFIIIC90 are transcriptional factor-related HAT families. In HATs, p600, SRC1, CLOCK, and AIB1/ACTR/SCR3, etc, several steroid receptor co-activators are included. In the cytoplasm, type B HATs are found that are responsible for acetylating the newly synthesized histones to their assembly into nucleosomes. HAT1 is an example of type B HAT members and their contribution to DNA repair and histone deposition [19].
The GNAT Family
The GANT family, Gcn5-related N-acetyltransferases is characterized by the presence of bromodomain and is located in the lysine residues on histones H2B, H3, and H4. The GNAT family members are marked by up to four conserved motifs (A-D) located in the catalytic HAT domain. The GNAT family comprises the most highly conserved motif A, and the sequence is Arg/Gln-X-X-Gly-X-Gly/Ala, which is important for the recognition and binding of acetyl-CoA (Loredana Verdone, 2005). In GNATs motif C was also found, but the majority of other known HATs are not present. The Gcn5 HAT is one of the best-characterized members in yeast. It includes four functional domains: N-terminal domain, c-terminal bromodomain, an Ada2 interaction domain, and a highly conserved catalytic (HAT) domain. In mammalian GNATs, the PCAF (p300/CBP-associated factor) and GCN5 are sharing a homology throughout their sequences.
N-terminal region of these proteins contain 400 residue which is absent in yeast Gcn5, but their HAT functions are preserved. The first HAT protein was Hat1 that is responsible for the cytoplasmic HAT activity in yeast and also binds strongly to histone H4. The example of type A HAT found in yeast is Elp3, which is part of the RNA polymerase II holoenzyme and plays an important role in the elongation of transcription [23].
The MYST HATs Family
Based on Four members MOZ, Ybf2 (Sas3), Sas2, and Tip60 name, the MYST family of HATs has been named. Esa1, MOF, MORF, and HBO1 are other important members. The presence of zinc fingers and chromodomains help to characterize these HATs families, which are located in the acetylated lysine residues on histones H2A, H3, and H4. Several MYST family proteins highly preserved motif A, which are important for acetyl-CoA binding. The N terminus region of the HAT domain of MYST proteins rich in cysteine is involved in zinc binding that is necessary for HAT activity. The first human MYST family member was Tip60 (Tat interactive protein, 60kDa) exhibit HAT activity. A homolog of MOZ (monocytic leukemia zinc finger protein) is Sas3 located in yeast, an oncogene located in humans. Esa1 was the first necessary HATs found in yeast. The HAT activity is very important for the transcription of the male X chromosome in flies. HBO1 (HAT bound to ORC1) was the first HAT Human, associated with the replication complex origin. MORF that are MOZ-related factors, shows close homology to MOZ throughout the entire length [3].
Histone Acetylation and Gene Regulation
Histone acetylation and deacetylation are necessary for regulation of gene. The acetylation was maintained by different specific promoters such as HATs and HDACs; explore their performed over large chromosomal domains denote that the acetylation state is in a continuous genome-wide flux. Histone acetylation and deacetylation regulate the transcription process by manipulating the higher-order folding properties of the chromatin fiber. Chromatin structure is not merely altered by acetylation of lysine residues but also give unique binding surfaces for repressors and transcription activators. Histone acetylation and regulation of transcription connect to similar molecular mechanism in yeast and multicellular eukaryotes but show a different ratio in between restrictive chromatin and permissive [23]. The role of histone acetylation in regulating gene expression is the main point. Acetylation of histone protein changes the nucleosome structure, or it provides protein binding signal. According to the changes in the nucleosomes structure, an acetyl group added to the lysine ε-amino groups of N-terminal tails and neutralizes the positive charge of the lysine side chain. This modification may affect the interaction between DNA and the lysine residue and contact become weaker with the negatively charged DNA backbone. According to the histone acetylation, it provides a signal for protein binding and added acetyl group to create a new surface for protein association. Docking sites are created by lysine acetylation for a protein molecule which called bromodomain.
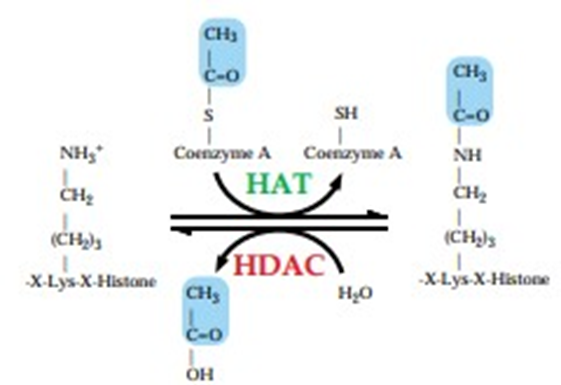
Figure 1: Histone Modification Due to Acetylation/Deacetylation [3]
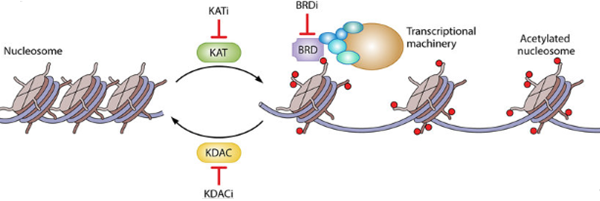
Figure 2: Regulation of Transcription by Histone Acetylation
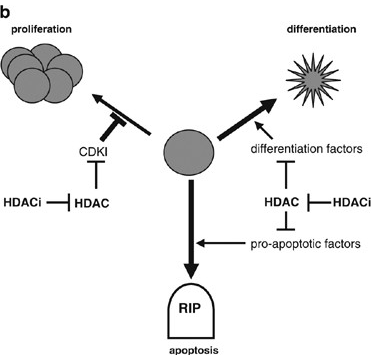
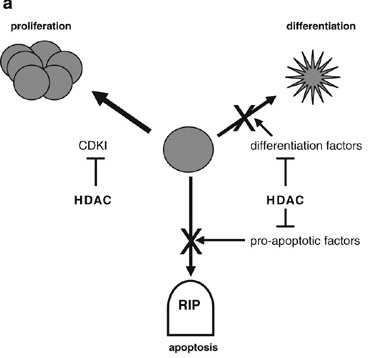
Figure 3: HDAC-Mediated Repression of Genes can Cause Uncontrolled Cell Growth. HDACs Repress The Transcription of (1) Cyclin-Dependent Kinase Inhibitors (CDKI), Allowing Continued Proliferation; (2) Differentiation Factors, Allowing Proliferation Instead of Differentiation; and Proapoptotic Factors, Permitting Survival. (b) HDAC Inhibitors (HDACi) can Restore Appropriate Gene Expression, Preventing the Uncontrolled Growth of the Cell. HDAC, Histone Deacetylases (MA Glozak, 2007) (v) Sirtuins Inhibitors: Nicotinamide (all Class III HDAC specific), Sirtinol (SIRT1, SIRT2 specific), Cambinol (SIRT1, SIRT2 specific), EX-527 (SIRT1, SIRT2 specific) [13,24]
This bromodomain located in the transcription and chromatin regulators. Different species carry different number like in human there are more than 60 bromodomain proteins, in Drosophila the number is 16 and in Saccharomyces cerevisiae 10 Bromo proteins present. But all bromodomains can’t recognize acetyl-lysine. It is reported that bromodomain containing protein, p300 binds to histone in acetylation. Chromatin acetylation by HATs is the function of bromodomain in gene regulation. In S. cerevisiae Gcn5 HAT bromodomain is needed for SAGA complex to associate with nucleosome remodeling and acetylated chromatin. Gcn5 bromodomain identifies Lys16-acetylated [23].
Another important yeast protein, Bdf1 has targeted the H4 histone tail. That protein has been needed for anchoring TFIID to the PHO5 promoter of S. cerevisiae, when the nucleosome relocating altered TATA which is H4 acetylation dependent manner.
Another important yeast protein, Bdf1 has targeted the H4 histone tail. That protein has been needed for anchoring TFIID to the PHO5 promoter of S. cerevisiae, when the nucleosome relocating altered TATA which is H4 acetylation dependent manner.
Bromodomain containing acetylated histone proteins was recently established in the living cells by FRET analysis. The regulator of transcription Brd2 connected with the acetylated lys12 of H4 and to a lesser degree with H2B but not with H2A and H4. Conversely, TFII250 is a component of TFIID, was found to recognize H4 and H3 that is of the lesser extent with H2B not H2A. PCAF, third bromodomain containing protein, was analyzed and produce signals of FRET with H4 and H3, not H2A or H2B.
Histone Deacetylase
Acetylation is a sort of epigenetic remodelling. of chromatin by which histone protein tightly wrapped around DNA is acetylated and relaxed. Due to this acetylation, the DNA becomes accessible for transcription factors, co-factors, RNA polymerase II & transcription proceeds. This acetylation is catalysed by the enzyme HAT. Another enzyme called HDAC counteracts the activity of HAT by removing the acetyl group from the core histone protein. The transition state of substrate is stabilized by tyrosine a histone residue that encourages the acetyl group to be released from histone lysine [24]. This is known as deacetylation & it reforms the compact structure of chromatin. In eukaryotes, HDACs can regulate transcription as well as gene expression by acetylation and deacetylation of the lysine residue of the histone proteins’ N-terminal domain. [3].
Classifications and Activity
HDACs are classified into four categories based on their resemblance to yeast proteins:
Class I: HDAC1, HDAC2, HDAC3 and HDAC8
Class II: (IIa) HDAC4, HDAC5, HDAC7, HDAC9 and (IIb) HDAC6, HDAC10
Class III: SIRT1 to 7
Class IV: HDAC11
Class I shows similarity to yeast RPD3, class II shows similarity to yeast HDAC1, class III shows similarity to yeast SIR2 & Class IV has features of sophistication class I & II HDACs [25-26]. Class I is found within the nucleus near the N-terminal portion. For HDAC1 the nuclear-localized sequence is encoded by the C-terminal located sequence KKAKRVKT and it must transfer to the nucleus [24]. Class II cells can traverse between the cytoplasm and the nucleus by phosphorylation suggesting the involvement of HDAC in non-histone protein acetylation. HDACs are actively engaged in deacetylation using two mechanisms depending on which they can be classified into two families: Zn+ dependent HDACs & NAD+ dependent HDACs. In Zn+ dependent pathway, the acetylated substrate present within the catalytic centre of the enzyme is stabilized by the Zn+ ion, and the carbonyl group in acetyl becomes polarized which facilitates the water molecule's nucleophilic interaction to the carbon atom and this is sensitive to TSA, SAHA. On the other hand, in NAD+ dependent pathway due to the transfer of acetyl group, nicotinamide and O-acetyl ADP ribose is formed and this is insensitive to TSA. [8,13].
Specificity of HDAC Activity
Class I HDACs in the nucleus may deacetylate all four subunits of histones, permitting transcription to proceed on the genome. The specificity of histone substrate for HDACs is quite difficult to be said because for two main reasons: Class I HDACs have the ability to deacetylate several lysine residues and the majority of Lysine residues can be deacetylated by multiple class I HDACs. Some nonhistone proteins such as p53, E2F1, the corepressor YY1, Proliferating Cell Nuclear Antigen (PCNA), and lysine demethylase1 are transcription factors due to the nuclear localization of HDAC1. These proteins show different effects on deacetylation such as p53 destabilized, E2F1 inactivated, YY1's repressive impact is decreased, as is PCNA's capability to bind to DNA polymerases beta and delta reduced, the ability of LSD1 to bind histone H3 enhanced. ERRα was shown to be deacetylated by HDAC8. As a result of this, the function of transcription factors is enhanced and the deacetylation of cohesin complex SMC3 encourages cohesin resynthesis after it has been removed from chromatin during anaphase or prophase. Class I HDACs also catalysed the post-translational modification by removing the acetyl group from the histone tail of H3 & H4.
In many multiprotein complexes, HDAC1 & HDAC2 are found together but in recent studies of humans and mice, it has been shown that HDAC2 can function independently from HDAC1 in vivo. The Myc, a transcriptional activator, and Mad, a transcriptional repressor, are both involved in the heterodimeric binding partner Max which is also a transcription factor. The DNA sequence containing the same E box binds to both Myc-Max and Mad-Max complexes but the HDAC1 and HDAC2 are required for mSin3 scaffold protein to be associate with the Mad-Max complex. Some co-repressor such as SMRT and N-CoR bind to HDAC3 effectively and this requirement of HDAC3 to promoters prevents the transcription. HDAC1 is required for repression of E2F-mediated transcription by Rb and this requirement leads to the decrease of acetylation at E2F-regulated promoter (MA Glozak, 2007). HDAC6 controls polyubiquitin-chain turnover and permits ubiquitylated proteins to be recognised and transported by binding its zinc-finger domain with ubiquitin. HDACs may be targeted to specified chromatin domains by noncoding RNA (ncRNA) (X-J Yang, 2007). Each protein has at least one catalytic domain where acetyl can bind and the localization of this domain is conserved within each class. The catalytic domain for the member of class I and class IV occupies most of the protein sequence. But the catalytic domain of class IIa HDACs always occupies the C-terminus of the protein sequence [24]. Class III HDAC SIRT2 prefers to deacetylate H4-K16 histone protein during the G2/M transition of cell cycle and it also influences the timing of S-phase entry [26].
A specific serine residue is phosphorylated which induces nuclear export of HDACs of class IIa and this triggers the 14-3-3 protein binding. This interaction can mask the NLS or unmask the NES. The short histidine residues which cannot reach the active site replace the catalytic Tyrosine 345 residue. This replacement of class IIa HDACs triggers the deacetylation of the acetylated histone without disturbing the transcriptional repression. Transcription factor RUNX2 is shown to be deacetylated by HDAC4. HDAC6 has two C-terminal zinc finger domains known as CD1 and CD2 which have a higher tendency for binding with free ubiquitin. It was shown that the CD1 prefers to deacetylate the C-terminal of the acetyl-lysine substrate whereas CD2 is effective on any acetyl-lysine substrate α-tubulin is deacetylated by HDAC6 which stabilizes the microtubules. Experiment has shown that the MSH2 is deacetylated and then ubiquitinated by HDAC6 which decreases its stability. HDAC11 can strongly show its activity for lysine defatty-acylation than deacetylation. It also deacetylates SHMT2 which influences the ubiquitination and degradation of proteasome of the IFN receptor [24].
HDAC can also deacetylate non-histone proteins, consequently modulating gene expression. HDAC1 from class I and SIRT1 from class III can deacetylate the tumour suppressor p53, HDAC3 from class I can deacetylate the p65 subunit of NF-kB which triggers its association with IkBa and leads to nuclear exportation of NF-kB. SIRT1 deacetylates the transcription factor p73 and SIRT3 deacetylates Ku70. This kind of deacetylation by HDAC can directly show effects on various physiologic processes, including inflammation, apoptosis, autophagy, metabolism, and differentiation [8]. Nonhistone proteins can be also affected by the HDAC present in the p300 which is a multiprotein complex [21].
HDACs have many other roles such as in metabolic processes, DNA metabolism by taking the acetyl group off of histone as well as nonhistone proteins. Some of the HDACs has also an important role in the replication of DNA, genomic stability, and the response to DNA damage for their pathogenic deregulation. Also in virology, it plays an important role. Experimentally it has shown that the human HDAC2 is the sole viral Non-structural protein 5 (NSP5) interactor in HEK293T cells SARS-CoV-2, which is responsible for the COVID-19 pandemic [24].
Disease Associated with HAT
Improper acetylation of a transcription factor may provoke diseases by imbalanced enzyme activities. The CBP/p300 HAT family is vital for differentiation, cell proliferation, apoptosis, and transformation. The occurrence of human chromosomal translocations further highlighted the CBP family of HATs, which is fuse CBP to MOZ (monocytic leukaemia zinc-finger protein), or MLL (mixed-lineage leukaemia). In humans these translocations can lead to leukaemia. In humans the p300 plays a tumour-suppressor role, has been investigated as well. Spontaneously arising mutations in the p300 gene have been a reason for various human cancers, as a result, two primary tumours and a numbers of cancer cell lines. In the human CBP gene mutations were related to Rubinstein-Taybi syndrome (RTS) [27] mental retardation, craniofacial defects, big toes, and broad thumbs are the reason for a developmental haploinsufficiency disorder. This disease is very common in humans, an average of 1 of 300 patients are suffering from mental retardation disease [12].
The imbalance of PCAF can stop growth. This is mainly happening due to the imbalanced interaction of PCAF and two important regulators of the cell cycle and these are E2F and p53. The PCAF indicates first the tumour suppression that comes from viral infection.
Tip60 also can relate with the interleukin-9 receptor and membrane receptor, and as a result, it is a reason for asthma and T cell oncogenesis [12].
The relation between the MOZ protein (monocytic leukaemia zinc finger protein) and cancer is better established. In leukaemia, several types of translocations have been observed. All fusions of different HATs are associated and interacted with two different HATs domains. So, this can postulate that the misdirection of HATs activity, either of the MOZ, the TIF2, or the CBP/p300 moiety is the reason for the malignant conversion type of leukaemia [28].
Disease Associated with HDAC
When deacetylation occurs in the normally active region and hyperacetylation occurs in the normal silence region, several diseases can develop. The activity of HAT/HDAC should be regulated, otherwise, dysregulation occurs that leads to various kinds of tumours, cancers, metabolic disorders, neurological disorders, and also the cardiological problem [12].
Cancer
A set of key genes which influence the growth of the cancer cells and plays an important role in their survival are strongly maintains by the HDACs. After knocking out the different HDACs (each was a member the of class I HDAC) from individual mice, the importance of HDACs in gene regulation was observed. The observation shows that the mice in which the HDAC1 was absent, died before birth due to proliferation defects. The mice in which HDAC2 was absent, died on the first day after birth due to cardiac malformation. The mice in which the HDAC3 was absent, died before birth due to gastrulation. According to the types of tumours, the levels of HDAC are varied in different types of cancer cells such as HDAC1 are present in prostate, lung, esophageal, colon and breast cancer cells; HDAC2 are present in colorectal, cervical and gastric cancer cells; HDAC3 is expressed in colon and breast tumour cells; HDAC6 are present in mammary tumour cells; HDAC8 is present in neuroblastoma cells and HDAC11 is expressed in rhabdomyosarcoma. Histone H4 hypoacetylation at lysine 16 linked with multiprotein complexes was highlighted as an epigenetic signature in cancer cells. The functions of some proteins which are most important in cancer (tumour protein p53, α3 subunit RUNX3, STAT3, β-catenin, estrogen receptor, EKLF, Myc, MyoD, NF-kB, HIF-1α, GATA family), are modulated by HDACs [13,26,29]. Acute myeloid leukaemia and urothelial are observed for low expression of HDAC5 but it can cause breast cancer with high expression. Renal cancer, non-small cell lung cancer, thyroid cancer is caused by the upregulation of HDAC9. Lung cancer and gastric cancer are caused by the low regulation of HDAC10 [24]. Due to the acetylation/deacetylation mechanism of many DNA repair proteins, the pathogenesis of tumours was elucidated which helps to discover the new drugs for post-translational modification for the treatment of cancer [25]. It is published in a recent study that tumour suppression can be influence by PIB5PA and its downregulation was also observed in melanoma. HDACs can cause histone hypoacetylation which leads to the downregulation of PIB5PA and decreases the survival of melanoma cells [10]. The inflammatory response is increased in an in vivo allergic inflammatory response model by HDAC1 and the parenchymal lung inflammation can also increase for HDAC1 deficiency in the Th2-type asthma model of mice [8].
Neurological Disease
Epigenetic changes that occur due to deacetylation by HDAC class I proteins are associated with neurological disease. It was observed from an experiment that the variation of Histone acetylation levels as a result of psychostimulant interaction are affected by HDACs [30]. Critical defects in brain development have been shown due to the deficiency of HDAC4. HDAC9 appears to be important for mature neuron function and deregulation of this can cause defects in brain cells [24]. Huntington's and Alzheimer's are two neurological diseases caused by the imbalance between histone acetylation and deacetylation [31-32].
Heart Disease
The deletion of HDAC3 in the heart causes interstitial fibrosis. In the T-cells in the interchange between intestinal lymphocytes and intestinal epithelial cells, the role of HDAC3 has also been observed. A loss of cardiac function has been shown after stress in HDAC5-null mice due to an increase in cardiac hypertrophy [24] In heart failure, HDACs have been discovered to interact with the transcription factor Mef2 and control gene expression [7].
Vascular Disease
The epigenetic modification of chromatin has a role in vascular diseases such as Vascular restenosis, atherosclerosis. Studies have indicated that HDAC activity can impact angiogenesis as a result of hypoxia, shear stress and VEGF-induced stem cell differentiation. It is evident that the HDAC7-null cell is involved in the development of normal vascular and in embryonic lethality [6].
HAT Inhibitors and its Role
Asthma, COPD, rheumatoid arthritis, and many cardiovascular diseases are caused by HATs. Hyperacetylation or acetylation pattern change both are connected in inflammatory diseases and restore the balance between acetylation and deacetylation will benefit those patients. Reduction of acetylation is a caused by HAT inhibitors. In the end, it leads to the responses of the different functions of HATs. Therefore, it is critical to identify which HATs are associated with inflammatory disorders and which require targeted to restore equilibrium. In viral diseases, HAT inhibitors can be used to treat patients. HATs enhance viral replication and viral protein invasion. HATs bind to viral proteins. E1A or Tat. It might be a promising target for inhibitor research. HATs play a crucial part in the control of transcription in Alzheimer's and Huntington's disease; however, it has been established that this is an excellent target for HAT inhibitors [33].
HAT subtypes have large differences in function but similarities in the structure that is might be vital caused of preventing side effects. Already, inhibitors are developed between selective HAT families, although selectivity of different functions and similar structures are very challenging. Generally, the natural products of HATs inhibitors are nonselective and inhibit a target besides HATs, including antioxidant activity, or protein reactivity. Some natural by-products have been developed between these selective HAT families [22].
The potential of Allosteric inhibitors to be selective between subtypes of the same family, because of the variation of N- and C-terminal regions, but the catalytic domain is identical for HATs families. These inhibitors are shown to exhibit biological effects, although in vitro it is still limited. HATs correlated with various proteins, modulating their activity but sometimes acetylation activity is not needed, like bromo domains or other protein interaction domains. Protein–protein interactions inhibition is the way of targeting specific HAT pathways [24].
HDAC Inhibitors
HDAC inhibitors are the types of compounds that play the opposite role of HDAC i.e., these proteins inhibit the deacetylation activity of HDACs. Due to its activity, it has a lot of therapeutic roles as well as it also helps in gene regulation. Sometimes even after preventing the condensation of the chromatin structure by inhibiting deacetylation, HDAC inhibitor does not always increase the transcription as this can have an impact on the function of transcription factors in both beneficial and negative ways.
Classifications
Based on their chemical structure, HDAC inhibitors may be classified into five types: (i) hydroxamates, (ii) short-chain fatty acids, (iii) benzamides, (iv) cyclic tetrapeptides and (v) sirtuin inhibitors. Based on their activity on HDAC, HDAC inhibitors can also be divided into two classes: HDAC specific inhibitors and HDAC-pan inhibitors.
Hydroxamates
Trichostatin A (HDAC-pan), SAHA (HDAC-pan), Belinostat (HDAC-pan), Panabiostat (HDAC-pan), Givinostat (HDAC-pan), Resminostat (HDAC-pan), Abexinostat (HDAC-pan), Quisinostat (HDAC-pan), Rocilinostat (HDAC-II specific), Practinostat (HDAC-I, II, IV specific), CHR-3996 (HDAC-I specific).
Short-Chain Fatty Acids
Valproic acid (HDAC-I, IIa specific), Butyric acid (HDAC-I, II specific), Phenylbutyric acid (HDAC-I, II specific).
Benzamides
Entinostat (HDAC-I specific), Tacedinaline (HDAC-I specific), Mocetinostat (HDAC-I, IV specific).
Cyclic Tetrapeptides
Romidepsin (HDAC-I specific).
Specificity and Role of HDAC Inhibitors
The different inhibitors can show a different anticancer effect on the different types of cancer. VPA can show its activity in bladder cancer but not in prostate cancer (Tomas Eckschlager, 2017). HDAC inhibitor of hydroxamic acid (hydroxamate) group can be specific as well as nonspecific. The activity in inhibition of HDAC enzyme depends on the ability of hydroxamic acid to form a chelate complex with Zn+ cation in the active site of enzyme HDAC. Pan-HDAC TSA inhibitor was used to treat hepatocarcinoma HepG2 cells. and the acetylation status of histone H3/H4 was observed. After observation, it was seen that the hyperacetylation of the TSS region appeared as transient but the TSS region downregulated by TSA inhibitor showed a strong reduction in acetylation over time [24].
Butyrate is a nonspecific HDAC inhibitor that has many biological effects such as cell cycle arrest, growth suppression, etc. The link between histone deacetylation and cell-static effect is strengthening by HDAC inhibitor Trichostatin A which induces differentiation [29]. It can also reduce cardiomyocyte hypertrophy and as a result of this, the non-muscle cells within the heart are affected which causes various inflammatory responses linked with toxicity. FDA approved the use of vorinostat and romidepsin HDAC inhibitors for the treatment of cutaneous T-cell lymphoma. HDAC inhibitors can also be used for the treatment of chronic arthritis and heart failure [7]. Romidepsin has its role in clinical treatment in which the cells in G2/M transition are blocked by increasing the tumour suppressor p19, p21 and p27 [24]. HDAC inhibitor SAHA plays a crucial role in the treatment of breast cancer, colon cancer, chronic lymphocytic leukaemia by inducing autophagy which again regulates the degradation of p53 mutant, further influence the survival of cancer cells. It also increases the radiosensitivity by targeting RAD51 and use for the treatment of pancreatic cancer. Some HDAC inhibitor such as rocilinostat, tubacin, tubastatin A etc are HDAC6 specific. HDAC6-specific inhibitors can block the function of the domain present in HDAC6 and due to this ability, it plays important role in bladder cancer, malignant melanoma, etc. Assisting the acetylation of α-tubulin by inhibiting HDAC6, they can regulate cancer cell cycle and apoptosis [34]. HDAC inhibitors can also inhibit the repressive effect of tumour suppressor p53. Some HDAC inhibitors can prevent the formation of the new vessel [26]. Valproic acid is used in the clinical trial for different types of tumours such as colorectal, breast, prostate, melanoma, NSCLC and pancreaticobiliary. Benzamide derivative compounds are also used for the clinical trial for leukaemia, pancreatic cancer, lymphoma, Hodgkin’s lymphoma, SHH medulloblastoma and other solid tumours. HDAC inhibitor in combined with PD-L1 antibody can show the antitumoral activity by modulating the gene regulation [24]. The activity of HDAC3 in HCT116 cells is inhibited by HDAC inhibitor SFN which promotes the degradation of HDAC3 and this facilitates the degradation and acetylation of CtIP by weakening the interaction between HDAC3 and CtIP. This modulates the HR activity in cancer treatment [25]. Some pan-HDAC inhibitor, as well as class-selective HDAC inhibitor, is used for the treatment of neurodegenerative disease [30]. A positive effect on memory and synaptic plasticity has been shown due to HDAC inhibitor and it also can be used as a mood stabilizer. Negative regulation of long-term memory is caused by HDAC3 [27].
The Significance of Histone Acetylation and Deacetylation in the Application of Biology
The chromatin decondensation due to the deacetylation in H3 and H4 lysine residues can control the transcription process and, also influence DNA repair, replication and cell differentiation [35].
The imbalance between acetylation and deacetylation can lead to proliferation and differentiation which causes several diseases and thus epigenetic modification can be used in therapeutic drug developments. SIRT1 was reported to use as an important tool in the treatment of cardiovascular disease. The damage caused by heart failure is protected by SIRT1 and SIRT3 [6,21]. HDAC inhibitors are widely used for a clinical trial to increase the activity of the drugs for the treatment of malignancies [13].
It has been shown that the innate immune cell at the transcription level can be controlled by acetylation. Acetylation occurs in the enzymes that are involved in the gluconeogenesis, glycolysis, TCA cycle, FAO, and the urea cycle. The presence of SIRTs in mitochondria is evident for the occurrence of the acetylation and deacetylation within the mitochondria. Acetylation in the mitochondrial proteins is responsible for several malfunctions as well as linked to the nutrient levels [9]. Histone modification can also be controlled by metabolites and thus the histone medication indirectly affects the metabolism. Some metabolic products such as acetyl-CoA, fumarate, SAM, succinate, 2-HG, NAM, NAD+, are used as the substrate in histone modification, controlling the metabolism process as well [2].
Acknowledgement
We would like to express our gratitude to Jadav University and Heritage Institute of Technology for proving us the opportunity and support to work on this review.






